-Search query
-Search result
Showing 1 - 50 of 59 items for (author: kawamoto & a)

EMDB-32892: 
Native Photosystem I of Chlamydomonas reinhardtii
Method: single particle / : Kurisu G, Gerle C, Mitsuoka K, Kawamoto A, Tanaka H

EMDB-32907: 
PSI-LHCI from Chlamydomonas reinhardtii with bound ferredoxin
Method: single particle / : Kurisu G, Gerle C, Mitsuoka K, Kawamoto A, Tanaka H

PDB-7wyi: 
Native Photosystem I of Chlamydomonas reinhardtii
Method: single particle / : Kurisu G, Gerle C, Mitsuoka K, Kawamoto A, Tanaka H

PDB-7wzn: 
PSI-LHCI from Chlamydomonas reinhardtii with bound ferredoxin
Method: single particle / : Kurisu G, Gerle C, Mitsuoka K, Kawamoto A, Tanaka H
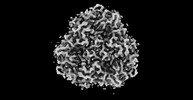
EMDB-32588: 
The Cryo-EM structure of siphonaxanthin chlorophyll a/b type light-harvesting complex II
Method: single particle / : Seki S, Nakaniwa T, Castro-Hartmann P, Sader K, Kawamoto A, Tanaka H, Qian P, Kurisu G, Fujii R

PDB-7wlm: 
The Cryo-EM structure of siphonaxanthin chlorophyll a/b type light-harvesting complex II
Method: single particle / : Seki S, Nakaniwa T, Castro-Hartmann P, Sader K, Kawamoto A, Tanaka H, Qian P, Kurisu G, Fujii R

EMDB-32041: 
Complex structure of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) pore with short stem
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

EMDB-32043: 
Complex structure of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) pore with long stem
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

EMDB-33188: 
Complex map of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) di-heptamer
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

EMDB-34136: 
Complex structure of Clostridioides difficile binary toxin folded CDTa-bound CDTb-pore (short).
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

EMDB-34137: 
Complex structure of Clostridioides difficile binary toxin unfolded CDTa-bound CDTb-pore (short).
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

PDB-7vnj: 
Complex structure of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) pore with short stem
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

PDB-7vnn: 
Complex structure of Clostridioides difficile enzymatic component (CDTa) and binding component (CDTb) pore with long stem
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

PDB-7yvq: 
Complex structure of Clostridioides difficile binary toxin folded CDTa-bound CDTb-pore (short).
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

PDB-7yvs: 
Complex structure of Clostridioides difficile binary toxin unfolded CDTa-bound CDTb-pore (short).
Method: single particle / : Yamada T, Kawamoto A, Yoshida T, Sato Y, Kato T, Tsuge H

EMDB-31605: 
Cryo-EM structure of cyanobacterial photosystem I in the presence of ferredoxin and cytochrome c6
Method: single particle / : Li J, Kurisu G

PDB-7fix: 
Cryo-EM structure of cyanobacterial photosystem I in the presence of ferredoxin and cytochrome c6
Method: single particle / : Li J, Kurisu G

EMDB-33554: 
Cryo-EM structure of MrgD-Gi complex with beta-alanine
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

EMDB-33555: 
Cryo-EM structure of apo-state MrgD-Gi complex (local)
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

EMDB-33556: 
Cryo-EM structure of MrgD-Gi complex with beta-alanine (local)
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

EMDB-33557: 
Cryo-EM structure of apo-state MrgD-Gi complex
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

PDB-7y12: 
Cryo-EM structure of MrgD-Gi complex with beta-alanine
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

PDB-7y13: 
Cryo-EM structure of apo-state MrgD-Gi complex (local)
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

PDB-7y14: 
Cryo-EM structure of MrgD-Gi complex with beta-alanine (local)
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

PDB-7y15: 
Cryo-EM structure of apo-state MrgD-Gi complex
Method: single particle / : Suzuki S, Iida M, Kawamoto A, Oshima A

EMDB-30353: 
Split conformation 2 of CtHsp104 (Hsp104 from Chaetomium Thermophilum)
Method: single particle / : Inoue Y, Hanazono Y, Noi K, Kawamoto A, Kimatsuka M, Harada R, Takeda K, Iwamasa N, Shibata K, Noguchi K, Shigeta Y, Namba K, Ogura T, Miki K, Shinohara K, Yohda M

EMDB-31520: 
Negative stain map of chained F1-like ATPase complex of Mycoplasma mobile
Method: single particle / : Toyonaga T, Miyata M

EMDB-30360: 
34-fold symmetry salmonella MS ring
Method: single particle / : Kawamoto A, Miyata T, Makino F, Kinoshita M, Minamino T, Imada K, Kato T, Namba K

EMDB-30361: 
11 fold-syymetry of salmonella MS ring
Method: single particle / : Kawamoto A, Miyata T, Makino F, Kinoshita M, Minamino T, Imada K, Kato T, Namba K

EMDB-30363: 
Without imposing symmetry of salmonella MS ring
Method: single particle / : Kawamoto A, Miyata T, Makino F, Kinoshita M, Minamino T, Imada K, Kato T, Namba K

EMDB-30613: 
Structure of the bacterial flagellar basal body
Method: single particle / : Kawamoto A, Miyata T, Makino F, Kinoshita M, Minamino T, Imada K, Kato T, Namba K

EMDB-30940: 
Salmonella flagella MS-ring protein FliF 1-503 having C33 symmetry
Method: single particle / : Makino F, Kawamoto A, Namba K

EMDB-30941: 
Salmonella flagella MS-ring protein FliF 1-456 having C35 symmetry
Method: single particle / : Makino F, Kawamoto A, Namba K

EMDB-30942: 
Salmonella flagella MS-ring protein FliF 1-456 having C34 symmetry
Method: single particle / : Makino F, Kawamoto A, Namba K
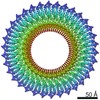
EMDB-30612: 
34-fold symmetry Salmonella S ring formed by full-length FliF
Method: single particle / : Kawamoto A, Miyata T

PDB-7d84: 
34-fold symmetry Salmonella S ring formed by full-length FliF
Method: single particle / : Kawamoto A, Miyata T, Makino F, Kinoshita M, Minamino T, Imada K, Kato T, Namba K

EMDB-30352: 
Split conformation 1 of CtHsp104 (Hsp104 from Chaetomium Thermophilum)
Method: single particle / : Inoue Y, Hanazono Y, Noi K, Kawamoto A, Kimatsuka M, Harada R, Takeda K, Iwamasa N, Shibata K, Noguchi K, Shigeta Y, Namba K, Ogura T, Miki K, Shinohara K, Yohda M

EMDB-30349: 
Staggered ring conformation of CtHsp104 (Hsp104 from Chaetomium Thermophilum)
Method: single particle / : Inoue Y, Hanazono Y, Noi K, Kawamoto A, Kimatsuka M, Harada R, Takeda K, Iwamasa N, Shibata K, Noguchi K, Shigeta Y, Namba K, Ogura T, Miki K, Shinohara K, Yohda M

PDB-7cg3: 
Staggered ring conformation of CtHsp104 (Hsp104 from Chaetomium Thermophilum)
Method: single particle / : Inoue Y, Hanazono Y, Noi K, Kawamoto A, Kimatsuka M, Harada R, Takeda K, Iwamasa N, Shibata K, Noguchi K, Shigeta Y, Namba K, Ogura T, Miki K, Shinohara K, Yohda M

EMDB-0977: 
Cyanobacterial PSI Monomer from T. elongatus by Single Particle CRYO-EM at 3.2 A Resolution
Method: single particle / : Kurisu G, Coruh O

PDB-6lu1: 
Cyanobacterial PSI Monomer from T. elongatus by Single Particle CRYO-EM at 3.2 A Resolution
Method: single particle / : Kurisu G, Coruh O, Tanaka H, Gerle C, Kawamoto A, Kato T, Namba K, Nowaczyk MM, Rogner M, Misumi Y, Frank A, Eithar EM

EMDB-30378: 
Salmonella FliF-FliG ring complex
Method: single particle / : Kawamoto A, Kato T, Kinoshita M, Minamino T, Namba K

EMDB-30379: 
11-fold symmetry of Salmonella FliF-FliG ring complex
Method: single particle / : Kawamoto A, Kato T, Kinoshita M, Minamino T, Namba K

EMDB-30233: 
Cryo-EM structure of the human pathogen Mycoplasma pneumoniae P1
Method: single particle / : Kawamoto A, Kenri T

PDB-7bwm: 
Cryo-EM structure of the human pathogen Mycoplasma pneumoniae P1
Method: single particle / : Kawamoto A, Kenri T, Namba K, Miyata M

EMDB-0713: 
Complex structure of Iota toxin enzymatic component (Ia) and binding component (Ib) pore with short stem
Method: single particle / : Yoshida T, Yamada T

EMDB-0720: 
Complex structure of Iota toxin enzymatic component (Ia) and binding component (Ib) pore with long stem
Method: single particle / : Yoshida T, Yamada T

EMDB-0721: 
Pore structure of Iota toxin binding component (Ib)
Method: single particle / : Yoshida T, Yamada T

PDB-6klo: 
Complex structure of Iota toxin enzymatic component (Ia) and binding component (Ib) pore with short stem
Method: single particle / : Yoshida T, Yamada T, Kawamoto A, Mitsuoka K, Iwasaki K, Tsuge H

PDB-6klw: 
Complex structure of Iota toxin enzymatic component (Ia) and binding component (Ib) pore with long stem
Method: single particle / : Yoshida T, Yamada T, Kawamoto A, Mitsuoka K, Iwasaki K, Tsuge H
Pages:
 Movie
Movie Controller
Controller Structure viewers
Structure viewers About EMN search
About EMN search



 wwPDB to switch to version 3 of the EMDB data model
wwPDB to switch to version 3 of the EMDB data model
